| parameters,[palette=None],[save=None]) |
>>> from pNbody import *
>>> nb = Nbody('gadget_z00.dat',ftype='gadget')
>>> nb.display()
>>> nb.display(size=(10000,10000),shape=(256,256),palette='lut2')
>>> params = {"size":(10000,10000),"shape":(256,256),"filter_name":'convol'}
>>> nb.display(params,palette='rainbow4')
| parameters) |
>>> from pNbody import *
>>> nb = Nbody('gadget_z00.dat',ftype='gadget')
>>> nb.show()
>>> nb.show(size=(10000,10000),shape=(256,256),palette='lut2')
>>> params = {"size":(10000,10000),"shape":(256,256),"filter_name":'convol'}
>>> nb.show(params,palette='rainbow4')
| parameters) |
It calls the methods CombiMap that create the matrix mat.
Then, it makes an integer matrix (matint) calling the function set_ranges and optionally add contours or box, using the functions contours and add_box.
It finally returns a tuple (mat,matint,mn_opts,mx_opts,cd_opts)
>>> from pNbody import *
>>> nb = Nbody('gadget_z00.dat',ftype='gadget')
>>> mat,matint,mn_opt,mx_opt,cd_opt = nb.Map()
>>> mplot(matint)
| parameters) |
This function returns a Float numarray matrix corresponding to the physical value defined by the parameter mode. The function uses either ComputeMeanMap, ComputeSigmaMap, or use directely the function ComputeMap to compute the desired matrix. The meaning of the parameter mode is given in the following table.
| Value | Meaning | Formula |
|---|---|---|
| "sr" | dispertion in r |
|
| "svr" | dispertion in vr |
|
| "svxyr" | dispertion in vxy |
|
| "svtr" | dispertion in vt |
|
| "szr" | ratio sigma z/sigma r |
If the value of mode may also be set to a variable or function name. In this case, a map using the function ComputeMeanMap, with a physical value taken as the given variable is returned.
>>> from pNbody import *
>>> nb = Nbody('gadget_z00.dat',ftype='gadget')
>>> nb = nb.select('gas')
>>> mat = nb.CombiMap(mode='T',filter_name='convol')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='log',cd=0,mn=0,mx=0)
>>> mplot(matint,palette='rainbow4')
>>> mat = nb.CombiMap(mode='log10(nb.T())',filter_name='convol')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='lin',cd=0,mn=7,mx=12)
>>> mplot(matint,palette='rainbow4')
| parameters,mode1) |
| (7.3) |
![]() is the zero momentum map and
is the zero momentum map and ![]() is the first momentum map of the variable given by mode1.
is the first momentum map of the variable given by mode1.
>>> from pNbody import *
>>> nb = Nbody('disk.dat',ftype='gadget')
>>> nb = nb.select('disk')
>>> mat = nb.ComputeMeanMap(filter_name='convol',size=(75,75),view='xy',mode1='z')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='lin',cd=0,mn=-10,mx=10)
>>> mplot(matint,palette='rainbow4')
| parameters,mode1,mode2) |
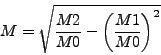 |
(7.4) |
![]() is the zero momentum map and
is the zero momentum map and ![]() (
(![]() ) is a first(second) momentum map of the variable given by
mode1(mode2).
) is a first(second) momentum map of the variable given by
mode1(mode2).
>>> from pNbody import *
>>> nb = Nbody('disk.dat',ftype='gadget')
>>> nb = nb.select('disk')
>>> mat = nb.ComputeSigmaMap(filter_name='convol',size=(75,75),view='xy',mode1='z',mode2='z2')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='lin',cd=0,mn=-2,mx=2)
>>> mplot(matint,palette='rainbow4')
| parameters) |
Compute the momentum map of an observable and return it in an float32 numarray matrix. It calls the functions :
The following table lists corresponding physical values fo the parameter mode that do not dependends on the observer position :
| Value | Meaning | Formula |
|---|---|---|
| "0" | zero momentum | |
| "m" | zero momentum | |
| "x" | first momentum in x | |
| "y" | first momentum in y | |
| "z" | first momentum in z | |
| "x2" | second momentum in x | |
| "y2" | second momentum in y | |
| "z2" | second momentum in z | |
| "vx" | first velocity momentum in x | |
| "vy" | first velocity momentum in y | |
| "vz" | first velocity momentum in z | |
| "vx2" | second velocity momentum in x | |
| "vy2" | second velocity momentum in y | |
| "vz2" | second velocity momentum in z | |
| "lx" | first specific kinetic momemtum in x | |
| "ly" | first specific kinetic momemtum in y | |
| "lz" | first specific kinetic momemtum in z | |
| "Lx" | first kinetic momemtum in x | |
| "Ly" | first kinetic momemtum in y | |
| "Lz" | first kinetic momemtum in z | |
| "u" | first momentum of specific energy | |
| "rho" | first momentum of density | |
| "T" | first momentum of temperature | |
| "A" | first momentum of entropy | |
| "P" | first momentum of pressure | |
| "Tcool" | first momentum of cooling time | |
| "Lum" | first momentum of luminosity | |
| "Ne" | first momentum of electronic density |
The following table lists corresponding physical values fo the parameter mode that do dependends on the observer position :
| Value | Meaning | Formula |
|---|---|---|
| "r" | first momentum of radial distance | |
| "r2" | second momentum of radial distance | |
| "vr" | first momentum of radial velocity | |
| "vr2" | second momemtum of radial velocity | |
| "vxyr" | first momentum of radial velocity in the plane |
|
| "vxyr2" | second momentum of radial velocity in the plane |
|
| "vtr" | first momentum of tangential velocity in the plane |
|
| "vtr2" | second momentum of tangential velocity in the plane |
|
>>> from pNbody import *
>>> nb = Nbody('disk.dat',ftype='gadget')
>>> nb = nb.select('disk')
>>> nb.cmcenter()
>>> nb.rotate(axis='x',angle=pi/4)
>>> mat = nb.ComputeMap(filter_name='convol',size=(50,50),mode='vr')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='lin',cd=0,mn=0,mx=0)
>>> mplot(matint,palette='rainbow4')
>>> mat = nb.ComputeMap(filter_name='convol',size=(50,50),mode='nb.rxyz()')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='lin',cd=0,mn=0,mx=0.0004)
>>> mplot(matint,palette='rainbow4')
>>> mat = nb.ComputeMeanMap(filter_name='convol',size=(50,50),mode1='nb.rxyz()')
>>> matint,mn_opt,mx_opt,cd_opt = set_ranges(mat,scale='lin',cd=0,mn=0,mx=50)
>>> mplot(matint,palette='rainbow4')
| obs,[eye=None],[dist_eye=None],[foc=None],[space='pos']) |
See About this document... for information on suggesting changes.